The forward primer creates copies of the 5’-3’ strand whereas the reverse primer makes copies of the complementary (runs 3’-5’) strand. 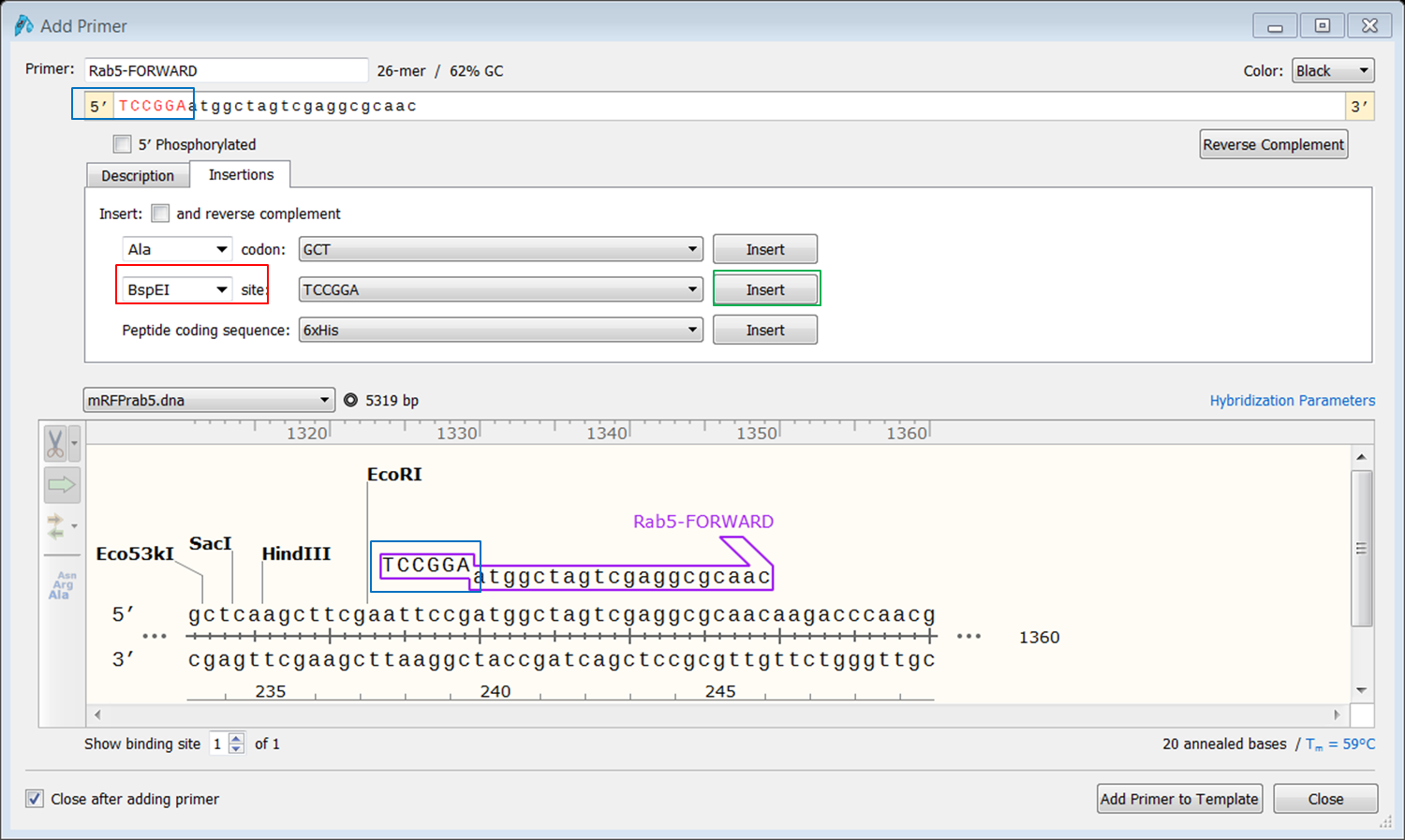
Whereas Reverse primer uses the upper strand as a template and synthesizes the lower strand.Īlthough the names suggest they create copies of different strands, their names depend on the direction of the strand being used for amplification. The forward primer synthesizes the upper strand using the bottom strand as a template. One is called ‘forward primer' and the other one is called ‘reverse primer’. To amplify any DNA sequence, two primers are necessary. Dsup protein is a newly discovered gene that imparts resistance to UV radiation by coating itself to DNA. Tardigrades are fascinating animals with extraordinary abilities to cope with extreme conditions like the vacuum of space, high tolerance for UV radiation, high and low-temperature tolerance to begin with. The primer design is demonstrated using Dsup (Damage Suppressor) gene from tardigrade (water bear). A User's Guide to the Arabidopsis T-DNA Insertional Mutant CollectionsĪ: Columbia-0 (CS60000, the sequenced genome)Ģ.This article demonstrates how to design primers (forward and reverse) for different types of cloning methods. PCR at the other end of the insertion? Is the right border less predictable in its insertion pattern?Ī: Please use the RB in the pBIN-pROK2 insertion sequences or try to use the sequences Q: Is there any way to design a Right Border primer for Q: Which T-DNA border flanking sequences wereĪmplified and sequenced? What are the primers do you use for left border PCR?Ī: The T-DNA left border sequence was usedįor PCR amplification of plant flanking sequences.įor PCR 1, we use LBa1 primer: 5' tggttcacgtagtgggccatcg 3'įor PCR 2 and sequencing, we use LBb1 primer: 5' gcgtggaccgcttgctgcaact 3'ģ. (it is actually transferred T-DNA) and our iSect primer design tool.Ĥ. Q: What is the number of T-DNA inserts perĪ: Approximately 50% of the lines containĪ single insert, the other 50% of lines contain two or more inserts.ĥ. Q: How can I obtain seeds for the Salk insertionĪ: Seeds for all sequenced indexed insertion Q: Are multiple T-DNA insertions amplifiedĪ: In most cases, we have identified only 1 T-DNA flankingĦ. Line are made available through ABRC and NASC. Seed requested should beĭirected to these stock centers. Q: The ABRC or NASC has sent me seeds for a Salk insertion Will not distribute seeds to individual laboratories.ħ. Q: What is the plant selectable marker used What generation (posttransformation) are these seeds?įrom ABRC and NASC are segregating T3 lines.Ĩ. Thus, it is not unusual for a mutant line However after several generations of growth, some of the lines OF BIOLOGICALLY-ACTIVE VIRAL SATELLITE RNA FROM THE NUCLEAR GENOME OF TRANSFORMED Q: What is the T-DNA transformation vectorīAULCOMBE DC, SAUNDERS GR, BEVAN MW, MAYO MA, HARRISON BD EXPRESSION To not express the drug resistance phenotype.ĩ. Of the pROK2 vector and the sequence of pBIN19.ġ0. Q: Which T-DNA border is used for plantĬontains an average of 1.5 -2.0 TDNA insertions. Due to the nature of the T-DNA integrationĮvent and the limitations of the PCR border recovery protocol, PCR productsĪre sequenced directly without separating the products of each reaction. It is up to the individual researcher to verify the Our BLAST search uses the best quality sequence to generate a single BLAST Thus, some sequencing reactions may contain two or more overlapping sequences. Left border of pROK2 and the position of the PCR amplification and sequencingĦ061 CGAGTGGTGATTTTGTGCCGAGCTGCCGGTCGGGGAGCTGTTGGCTGGCTGGTGGCAGGAĦ121 TATATTGTGGTGTAAACAAATTGACGCTTAGACAACTTAATAACACATTGCGGACGTTTT The left border of pROK2 and where do the PCR isolation and DNA sequencing The arrow point in the 5' to 3'ĭirection beginning with the left border DNA and into the plant flanking What are the arrows above the T-DNA line?Ī: The arrow indicates the orientation of Sequences of each PCR product from any line. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
